Samtools get consensus sequences9/12/2023  
ViReflow extracts the core steps of our production pipeline ( ), implemented directly into AWS, which we have used to process tens of thousands of sequences in UC San Diego’s Return to Learn program ( ) 12. ViReflow also implements optional analyses for specific viruses of interest, such as viral lineage calling (e.g. ViReflow implements the following standard viral consensus sequence workflow: (1) read trimming, (2) read mapping, (3) variant calling, and (4) consensus-sequence calling. ViReflow was developed specifically in response to the COVID-19 pandemic, but it is general to any viral pathogen. The ViReflow pipeline was built around Reflow, an incremental cloud-based data processing system developed by GRAIL ( ). ViReflow’s simplicity and ease-of-use is critical to adoption by public health professionals who may have limited experience with command line interfaces. In addition to being the only pipeline that supports viral lineage assignment 9 beyond just Pangolin 10 (via VirStrain 11), the key benefits of ViReflow over the existing tools are its automatic cloud compute resource scaling for rapid cost-optimized parallel processing and its intuitive GUI. A comprehensive pipeline comparison can be found in Table 1. To our knowledge, the only existing tools with similar functionality to ViReflow are V-pipe 5, the nf-core/viralrecon pipeline 6, HAVoC 7, and ViralFlow 8. Reflow was chosen for its ability to automatically dynamically scale resource allocations on AWS without intervention from the user. 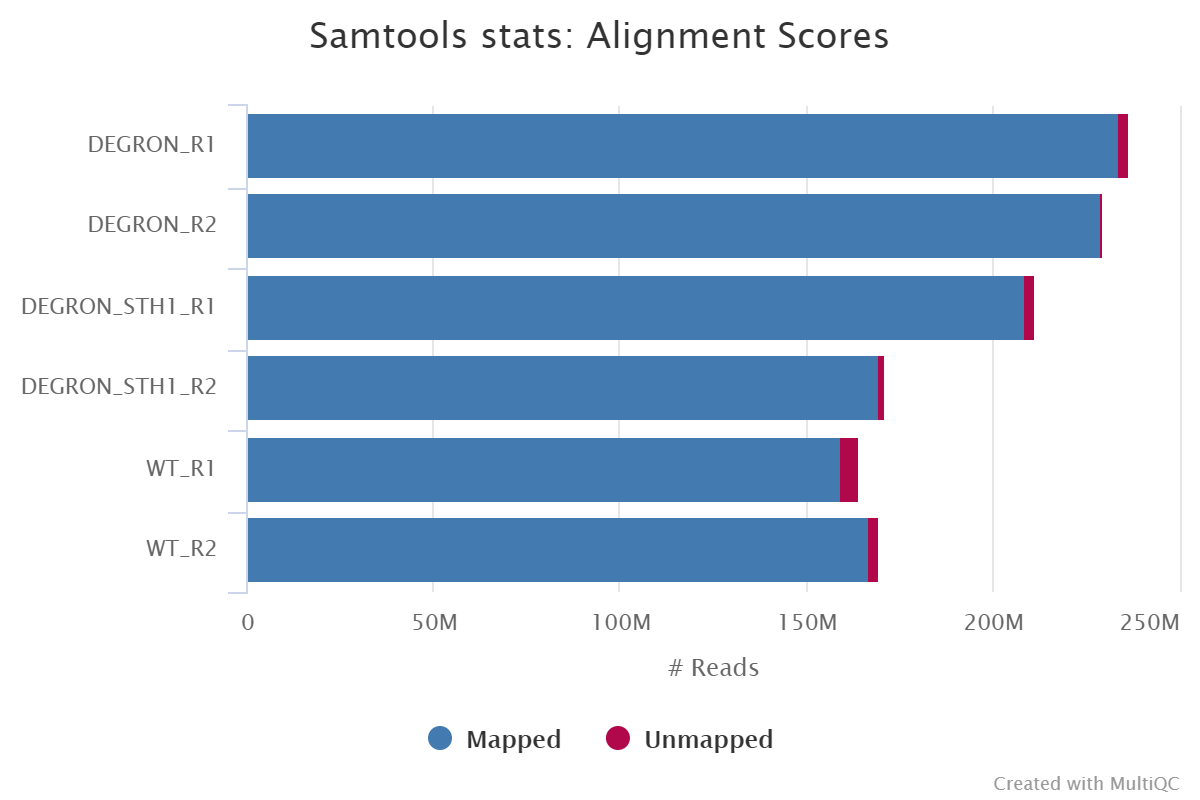
In this article, we present ViReflow, a user-friendly viral consensus sequence reconstruction and analysis pipeline enabling rapid analysis of large-scale viral sequence datasets using AWS and the Reflow system 4. Cloud computing platforms such as Amazon Web Services (AWS) are accessible and relatively inexpensive, but the optimal design and configuration of a cloud compute cluster typically requires systems administration expertise, and suboptimal cloud compute configuration can result in delays in time-to-results as well as in excess compute costs. Many labs have access to sequencing technologies, but relatively few have experience with high-performance computing resources. However, the sheer magnitude of raw viral sequence data that is collected poses a significant computational challenge. In a rapidly-growing pandemic, the time from raw sequence data to results (i.e., high-confidence variant calls and consensus sequences) is of utmost importance to implementing public health interventions in real-time. 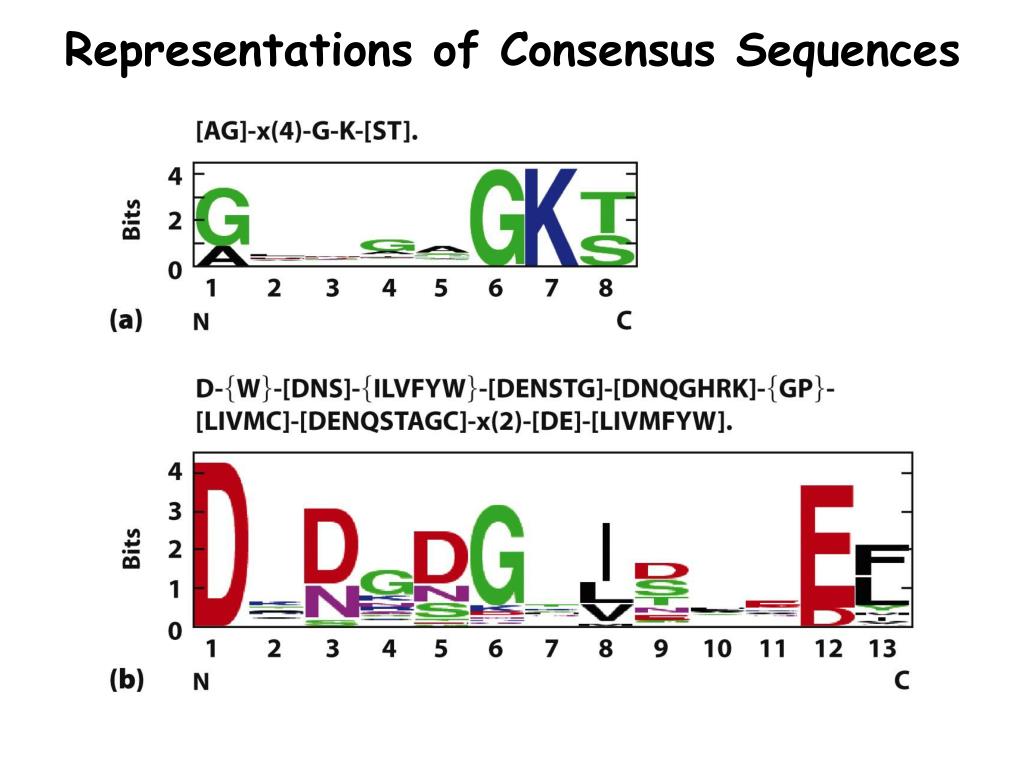
Improved throughput of and access to sequencing technologies has dramatically increased viral sequence data production: one sequencing run on an Illumina NovaSeq S4 flow cell can yield raw viral sequence data from > 1500 patient samples 2, and as of October 2021, over 4 million complete SARS-CoV-2 genomes have been deposited to the Global Initiative on Sharing All Influenza Data (GISAID) EpiCoV database 3. Molecular epidemiology uses viral genome sequences from patient samples to provide real-world public health insights about outbreaks 1. ViReflow’s simplicity, flexibility, and scalability make it an ideal tool for viral molecular epidemiological efforts. Importantly, when utilized with sufficient compute resources, ViReflow can trim, map, call variants, and call consensus sequences from amplicon sequence data from 1000 SARS-CoV-2 samples at 1000X depth in < 10 min, with no user intervention. We developed ViReflow, a user-friendly viral consensus sequence reconstruction pipeline enabling rapid analysis of viral sequence datasets leveraging Amazon Web Services (AWS) cloud compute resources and the Reflow system. However, with great sequencing comes great computation, and while cloud computing platforms bring high-performance computing directly into the hands of all who seek it, optimal design and configuration of a cloud compute cluster requires significant system administration expertise. Throughout the COVID-19 pandemic, massive sequencing and data sharing efforts enabled the real-time surveillance of novel SARS-CoV-2 strains throughout the world, the results of which provided public health officials with actionable information to prevent the spread of the virus. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
